The Laboratory of Computational Biology
Richard Pastor and Bernard Brooks
The Laboratory of Computational Biology is an interdisciplinary group of scientists who study biological processes via computer simulation. It is part of the Biochemistry and Biophysics Center Division of Intramural Research National Heart, Lung, and Blood Institute in the National Institutes of Health. The laboratory is a consortium of the laboratories lead by Dr. Bernard R. Brooks and Dr. Richard Pastor.

Groups
The Laboratory is consortium of two Labs and one section in one facility. Each laboratory is led by a principal investigator. A brief overview of their work is given here, and you can find out more in the web pages for each Lab.
Laboratory of Computational Biophysics
Dr. Bernard Brooks
The Laboratory of Computational Biophysics consists of researchers who both develop simulation and modeling techniques and apply them towards the study of problems of biological significance. Techniques employed include; molecular dynamics, quantum and molecular mechanics, ab initio analysis of small molecule structures, molecular modeling, and electron microscopy image analysis.
Laboratory of Membrane Biophysics
Dr. Richard Pastor
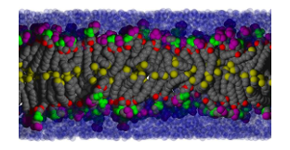
The Laboratory of Membrane Biophysics is a group of researchers who use diverse theoretical methods combined with high performance computing to investigate the properties of lipid membranes and related molecular assemblies. The Lab operates under the direction of Dr. Richard Pastor; the group joined the NIH/NHLBI Laboratory of Computational Biology (LCB) in 2006, after 20 years in FDA/CBER's Laboratory of Biophysics.
High Performance Computing Section
Dr. Daniel R. Roe

The High Performance Computing Section (HPCS) maintains and supports all the infrastructure of hardware and software of the LoBoS cluster that allow researchers of the laboratory to use high performance computing to investigate biological processes. It is also responsible for developing new software nd evaluating the suitability of emerging technological platforms for use in computational simulations. The section operates under the direction of Dr. Daniel R. Roe.

